Typically, access is provided across an institutional network to a range of IP addresses. If you are a member of an institution with an active account, you may be able to access content in one of the following ways: Get help with access Institutional accessĪccess to content on Oxford Academic is often provided through institutional subscriptions and purchases. To the best of our knowledge, HEAP will be a valuable tool for insight into the complex mechanisms of enhancer activity. 
Notably, the explainable framework HEAP utilizes post-hoc interpretation to provide insights into the prediction mechanisms from three perspectives: data, model architecture and algorithm, leading to a better understanding of model decisions and enhancer grammar. Experiments demonstrate that HEAP outperforms published methods and showcases the effectiveness of the TAPS, especially for those with limited training samples. Then we adapt to specific tasks by adding several task-specific subset layers. We first train a shared model with all cell-type datasets. We use a novel two-step multi-task learning method, task adaptive parameter sharing (TAPS), to efficiently predict enhancers in different cell types.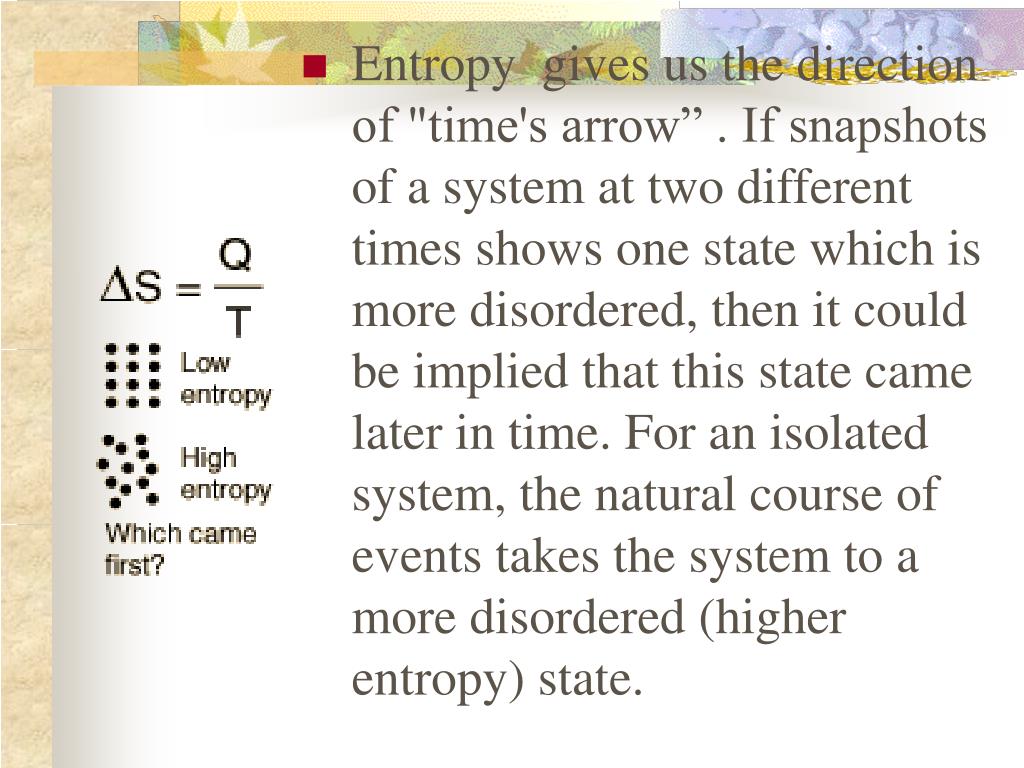 
The algorithm can incorporate DNA sequences and epigenetic modifications to obtain better accuracy. The framework includes three modules that use grammar-based reasoning for enhancer prediction. Here, we present HEAP ( high-resolution enhancer activity prediction), an explainable deep learning framework for predicting enhancers and exploring enhancer grammar. Despite extensive genetic and computational studies, accurately predicting enhancer activity in different cell types remains a challenge, and the grammar of enhancers is still poorly understood. Enhancers are crucial cis-regulatory elements that control gene expression in a cell-type-specific manner. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
